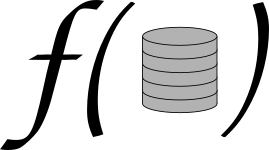
As researchers, our job is to develop new knowledge and share it with the world. Currently, the main way to do this is research articles. Thus, we are all contributing to a vast corpus of scholarly literature, which contains millions of papers across decades of research in all domains of human knowledge.
The point of this article is to argue that this precious resource is not currently used to its full potential. The scientific corpus can only be navigated with imperfect search engines, it is spread across multiple publishers, and it is often hidden behind paywalls. Thus, many innovative uses of this corpus are not possible yet. Compare this to the corpus of Wikipedia articles, which is much more high-level, but is easily and freely downloadable: the Wikipedia corpus has thus been used for information extraction, language model learning, integration with other knowledge graphs, and more. With the scientific corpus, all of this is mostly uncharted territory.
In this article, we will first present the new potential applications that we envision for the scientific corpus. Second, we will present the obstacles that prevent us from getting there, and how we could overcome them. We hope that this article can inspire our communities and generate some discussion about how to improve our current publication system: we have also created a mailing-list to discuss these questions further (more info at the end of the article).
What should be possible?
Better academic search engines
An important part of our job is to search for related work and other sources of inspiration. This is typically done using a search engine for scientific papers, such as:
- Google Scholar
- Microsoft Academic Live Search
- Subscription-based tools run by publishers, e.g., Web of Science, Scopus, MathSciNet
These solutions are widely used, but they are not ideal. Tools run by publishers can only be accessed with a costly subscription, and the Google and Microsoft search engines are only side projects of these large companies: they do not seem to receive much attention in terms of development, and could be discontinued at any time. There is ample room for improvement in this area by making better use of the scientific corpus, especially by innovative actors such as start-ups and research labs that are more invested in the area. One could imagine:
- Better recommendations, especially explainable recommendations. Google Scholar produces recommendations which are sometimes good, sometimes not, but always opaque. These algorithms can improve via user feedback, i.e., using the dataset indicating which researcher clicked on which recommendation, but we do not know whether Google does this, and in any case this dataset remains secret.
- Subscriptions to individual researchers, to communities, to themes. Google Scholar makes it possible to follow a researcher, but not to follow papers from this reearcher about a specific topic. It also does not know about communities, e.g., following papers published by the PODS and ICDT communities.
- Better fulltext search. Google Scholar indexes the full text of many research articles (via commercial partnerships with publishers), but not all of them.
- Better links between cited articles. Google Scholar can find articles citing a given article, and search within these articles. However, its citation information is often noisy, and cannot be corrected by users. Meanwhile, there are more modern initiatives to extract the citation data between papers and allow users to fix problems, e.g., the CROCI index, which are independent of Google Scholar.
- Open APIs. Google Scholar has no API, whereas Microsoft proposes the Academic Knowledge API. An academic search engine with a powerful API makes it possible to integrate information in other websites, and also other software, e.g., LaTeX editors. Wouldn’t it be nice if you could search for a paper directly in your text editor and automatically insert the corresponding BibTeX entry? Or add hyperlinks in article bibliographies, or find citing articles or errata, directly within your PDF reader?
Why have these ideas not inspired new players to emerge in the field of academic search engines? One big reason is that the barrier to entry is hard — as we will explain, the academic corpus is difficult to access.
Information extraction in articles
Going beyond existing search engine features, a much more ambitious goal would be to extract structured and semantic information from research articles. This would make it possible to find articles based on structured queries rather than textual search, e.g., searching based on mathematical formulas, complexity classes, graph classes, mathematical objects, etc. Extraction approaches could also understand the internal structure of an article (sections, theorems, figures, tables, etc.), to make it easy to jump to the right place. This is an ambitious information extraction task, but would unlock many new applications:
- Finding relationships between mathematical results. What are the individual results on which a given result depends? If you generalize a given result, or if you falsify it, then what could be impacted? What has been proven conditionally to another results, which results become true if you prove a given conjecture? Are there results in the literature that seem to contradict each other?
- Semantic citations. When a paper cites another paper, which precise part is being cited? (e.g., can I jump directly to “Theorem 4.2 of [13]”?) What is the meaning of the citation? (is it about using a result of the cited paper, extending it, modifying its proof, refuting it, applying it to a use case, presenting a variant of it, etc.)
- Hyperlinks. Can I jump directly, in an article, from a result to its proof? From an obscure name to the definition of the corresponding concept? From an arcane notation to the place where it was introduced?
- Disambiguation. Sometimes, different areas use the same name for different things: a “graph” can be directed or undirected, it can be a simple graph or it can allow multi-edges. Sometimes, different areas use different names to refer to the same thing (e.g., existential rules and TGDs). A smart search engine could figure this out to do more efficient searches and find related work from neighboring areas with a different terminology.
- Domain-specific inference. Could we develop a search engine that understands basic notions of complexity, to answer queries like “Which restrictions of 3-SAT remain NP-hard?” or “On which graph classes is 3-coloring tractable?”. Or answer queries about asymptotic complexity, like “What is the best algorithm to find squares in an input string? To find triangles in an input graph?” Or about graph classes, e.g., “Do bounded treewidth graphs also have bounded mim-width?” Or about TGD classes, query classes, Datalog classes, etc.
There are ad-hoc initiatives of this kind in several domains. For instance, in CS, we have the Complexity zoo for complexity classes, the ISGCI for graph classes, the House of graphs (and other resources) for interesting graphs, the Description logic complexity navigator, the OEIS for integer sequences, and the Open Problem Garden for open problems. But we could use the scholarly corpus to extend automatically these resources, connect them to the relevant literature, and create such tools for other domains in a more automatic fashion.
Textual data
Even without semantic inference, the raw textual data of scientific articles could be useful for many applications. For instance:
- Topic detection. Given an article, how to classify it in the correct field? How to find plausible reviewers for it? All these tasks could be solved automatically by training classifiers on the existing research corpus. This is explored, e.g., by the Toronto Paper Matching System, but only for the articles that it can access.
- Plagiarism detection. There are dedicated solutions for this, but they are not free. If the scientific corpus were publicly available, this would be much easier.
- Training a language model. For instance, modern AI techniques can automatically generate convincing sounding text: could they also generate plausible scientific articles? And which reviewers would they fool? :)
- Analysing trends. Which topics, keywords, etc., have become more or less fashionable over time? Imagine the Google Ngram Viewer over the computer science research corpus
Safekeeping and availability
If we could easily get copies of the scholarly corpus, it would also make it easier to backup and ensure that it is available over the long term. Currently, we trust each publisher to ensure the digital preservation of their articles, e.g., ACM, or arXiv. However, third parties cannot also easily do their own backup of articles, which would help create even more archive copies of our hard work.
Another use of such an archive would be offline reading: we still spend some time without good Internet access, so it is often convenient to save offline copies of the papers we work with. Wouldn’t it be convenient to directly download an indexed copy of all papers that we are likely to need, e.g., all papers ever published in database theory and neighboring areas? These would comfortably fit on a normal hard drive. The problem is to access these articles, as we will explain.
How could we make this possible?
We have presented many innovative ways to use our research work, and there would be lots of research to be done to make these innovative uses possible. However, to explore them, the first step is to prepare a dataset of the scientific corpus, in the right format.
First challenge: Making sense of PDF articles
The scientific literature corpus is largely made of PDF documents, formatted according to rigid and paginated layouts. This is good for printing, but offers little in terms of structured content: sections, figures, tables, theorems, references, and even lines and paragraphs are not clearly marked in the PDF file.
Fortunately, it is sometimes enough to work with the metadata of scientific articles, without looking at the content inside the PDF. To access this metadata, one useful tool is DBLP: they offer dumps and an API to retrieve bibliographic information (authors, titles, venues) for a large subset of the computer science literature (about 5M entries). Beyond computer science https://www.crossref.org/ provides an API with about 100M entries across all academic fields; BASE has information about around 100M open-access articles, and a corresponding API; OpenCitations has structured information about the references included in hundreds of thousands of recent research articles, accessible for download and through a SPARQL endpoint. S2ORC has structured information about about 80M papers, including around 380M citation edges.
Another direction is to use information extraction techniques to extract structured information from the PDF itself. Scholarly information extraction is an active area of research, with numerous freely available tools. Current systems, such as GROBID, can extract information such as title, authors, institutions, abstracts and lists of citations with reasonably high precision. A few other works, more experimental in nature, have looked into the extraction of other parts of a PDF article, such as figures and tables.
In some cases, in addition to the PDF article itself, the LaTeX article source is also available. For instance, arXiv offers the sources of most of their LaTeX-based articles. Extracting structured content from LaTeX sources is simpler than from PDF articles, though by no means trivial given that TeX is Turing-complete!
Finally, the authors or publishers of scientific articles can also manually add semantic annotations to their files. This is commonly done for the title and author list, for keywords and categories, but can also be done for much more. For instance, https://ctan.org/pkg/stex is a collection of LaTeX packages, under active development, that can express rich semantic information about mathematical formulas, mathematical statements, proofs, etc. It is for example possible to explicitly declare variables, to produce https://www.openmath.org/ mathematical formulas, to explicitly link a theorem to its proof, to structure a proof in different steps, to add some semantics to references and citations, etc.
In summary, it is complicated to extract information from papers in the scientific corpus because they are in PDF, but this can be worked around in several ways: working at the level of metadata, or performing information extraction from the LaTeX source or from the PDF. In the longer term, we can hope that more and more authors and publishers will add semantic markup to their articles, that more and more publishers will distribute the sources of articles like arXiv does, and that more steps will be taken to ensure that the scientific corpus is machine readable.
Second challenge: Obtaining the papers
However, before analysing the papers, the first step is to retrieve all of them. Sadly, this is already a complicated endeavor. On the face of it, one problem is that the scientific corpus is not centralized but hosted by many different publishers. However, this should not be an obstacle, as we could use the existing centralized metadata sources (Crossref, DBLP, etc.) to list all papers, and then download all of them. The size of the dataset is also not a huge obstacle: estimating that there are around 100M scientific articles (from Crossref) and that their average size is less than 1 MB, we get an upper bound of 100 TB of data. This is large, but could fit on a dozen of hard drives, requiring at most a few thousand euros of hardware.
The real issue is that many scientific papers are only available behind paywalls and can only be accessed from institutions having paid for the adequate subscriptions. Even with a subscription, the massive download of scientific articles is often blocked by technical measures and prohibited by the subscription conditions: see, for instance, the prosecution of Aaron Swartz in the US for massive download of JSTOR. Some publishers allow text and data mining on their corpus, e.g., Elsevier, but these solutions are local to a publisher, and they are limited and far less powerful than downloading directly the full dataset of research articles.
There are limited ways to work around paywalls. For instance, many recent scientific articles are available for free in open-access archives or on author webpages. However, it is challenging to link open-access material (e.g., found in BASE or crawled on the Web) to the bibliographic entries and DOIs to which they correspond: this task is explored by the Unpaywall project, and open-access links via Unpaywall have very recently been added to DBLP. Authors can encourage this approach by making all their articles freely available in open repositories, and annotating them with information connecting them to the publisher version, e.g., by adding DOI information to their arXiv deposits.
However, in addition to these, a better but more ambitious solution would be to ensure that scientific articles are made freely available on the Web, directly by scientific publishers. This is the open access movement. Indeed, publishers have little involvement in the research or writing of scientific articles, and the Internet has drastically reduced the costs of distributing articles, so it does seem odd that publishers should make money selling our work. In our community, ICDT moved in 2014 from paywalled ACM proceedings to open-access proceedings now hosted by LIPIcs. As far as PODS proceedings are concerned, we recently agreed with ACM to have them freely downloadable via their OpenTOC service, but this is only possible when accessing PODS articles through a specific URL on the PODS pages; regular access through the ACM digital library is still paywalled. In the computer vision community, for instance, the IEEE ICCV and CVPR conferences are using a hybrid model where official proceedings on IEEE Xplore cohabit with open-access proceedings provided by the Computer Vision Foundation. However, the PODS and ICCV/CVPR models do not solve licensing issues (which we describe in the next point). We track the status of various other conferences and journals relevant to the database community on this page.
Currently, the most effective way to retrieve scientific articles in practice are still pirate solutions. The most notable ones are the Sci-Hub portal which allows users to circumvent paywalls and serves around 400K requests per day, and the Library Genesis project which has compiled an archive containing tens of millions of scientific articles. The problem is that these efforts do not have clear legal standing, so they cannot attract the visibility and funding that would correspond to their considerable usefulness to the scientific community. Such datasets of papers have neveretheless being used to some extent for new applications such as text mining.
In summary, the first difficulty in preparing a dataset of the scientific corpus is to retrieve scientific articles, and the obstacle is not technical but legal. The best solutions do not currently have legal standing, and legal solutions will remain complicated as long as scientists will allow publishers to sell their work.
Third obstacle: Distributing the dataset
Assume now that we have managed to retrieve a large dataset of papers. The last challenge, and maybe the hardest one, is to be able to redistribute this dataset so that it can be used by other parties. Again, the challenge is not technical, but legal. You cannot legally redistribute papers that you obtained illegally, and even legally obtained papers cannot be redistributed in most cases. Papers obtained via a subscription cannot of course be redistributed freely, and even open-access papers obtained on repositories usually cannot be redistributed either. Indeed, closed-access publishers require authors to sign an exclusive copyright transfer: authors keep a right to post the articles in open repositories under some conditions, but they cannot give permission to others to redistribute the articles. Of course, publishers do this to minimize the impact of free redistribution on their profit margins.
For this reason, it would not be legal to harvest papers (via subscriptions or on free archives) and redistribute the resulting dataset. Likewise, the dataset of arXiv papers cannot be freely redistributed (which limits its usefulness, given that it cannot be downloaded for free either). Update: there is actually a mirror of the arXiv PDF files and sources on Archive.org, which is free to download; however, its copyright status is not completely clear.
The right solution to this problem is the one adopted by most open-access publishers: not only do they distribute their articles freely, but they distribute them under a Creative Commons license that permits free redistribution by anyone (under some conditions). These Creative Commons licenses are also used by Wikipedia, by Stack Exchange websites (e.g., Stack Overflow), and allow everyone to download and redistribute dumps of Wikipedia and Stack Exchange.
When scientific publishers use these permissive licenses, there is no obstacle to redistributing a corpus of scientific articles. To illustrate this, we have prepared a dataset of all scientific papers from LIPIcs (as of early 2019), i.e., 9404 papers that we have collected and can freely redistribute without even needing to ask LIPIcs or the authors for permission. This dataset contains in particular all ICDT papers published with LIPIcs over the last years. We hope that this dataset can be useful for many tasks: text analysis, similarity analysis, or any other innovative uses. If you build things out of this dataset, please redistribute them too (it should be legal in almost all cases)!
So, why can’t we have a dataset like this, but with a hundred million scientific papers instead of ten thousand? The main problem is the legal issue of closed-access publishing and exclusive copyright transfer, and we as a community are the ones responsible for it: because we continue to publish in these venues, to review for them, and to host them. Solutions are possible, in particular for PODS. For example, the ACM conference POPL has successfully transitioned to an open-access model; a similar model could probably be adopted for PODS if the community wanted it.
Call for interest
If you have read this far, you are probably interested in how to modernize and improve scholarly publishing in computer science. In this case, you are not alone! We would like to start a mailing-list of people who would like to push the system in the right direction, to better coordinate our efforts. If you would like to join us, you can subscribe here.
Authors
-
Antoine Amarilli, Télécom Paris, IP Paris
-
Pierre Senellart, École normale supérieure, PSL University
This article expresses the personal point of view of the authors and its contents are not endorsed by the author’s current or past employers.